- Blog
- About
- Contact
- Soul calibur 5 pc download mega
- Alpha zawgyi keyboard for window 7 64 bit
- Kana kanum kalangal online
- Unertl 8x usmc sniper scope
- Snyncing endnote with chrome
- Taylor swift today was a fairytale chords
- Vidmate app problems
- Stat transfer 10 download
- Trimble terramodel
- Hitfilm pro mac torrent
- Nitro pro 10 activation code
- High tail hall flash game
- Sim girl version 6-6 wiki
- Mmpi 2 scales for dummies
- Riddle school 3 button
- Protein oligomers hydrophobic amino acids
- Mikrotik routeros 6-28
- Enigma album
- What to do with psp iso files
- Eve aio bot not working after update
- Why are psp iso files so huge
- Acronis true image 2014 tutorial
- Manowar warriors of the world
- Keep2share premium link generator mobile
- Dum dum song lyrics
- Euro truck simulator 1 works windows 8
- A law abiding citizen
- Fastboot flash recovery twrp img
- Neon drive achievements
The formation of protein oligomer, known as protein assembly, is also a common reaction used by pathogens to produce killing “machineries”. In addition, numerous monomeric proteins associate transiently in binary or in higher stœchiometries (number of chains associated in a protein oligomer) during their life span. They are referred to as protein oligomers and have what is called a quaternary structure. Most proteins are made of more than one polypeptide chain to carry out their biological function. Supported by the University of Savoie (4000 euros)The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.įunding: This work was supported by The system complex Rhone Alpes IXXI (5000 euros). Received: OctoAccepted: JanuPublished: April 9, 2012Ĭopyright: © 2012 Feverati et al. Gisou van der Goot, Ecole Polytechnique Federale de Lausanne, Switzerland Such property would be extremely valuable in term of assembly inhibitory drug development.Ĭitation: Feverati G, Achoch M, Zrimi J, Vuillon L, Lesieur C (2012) Beta-Strand Interfaces of Non-Dimeric Protein Oligomers Are Characterized by Scattered Charged Residue Patterns. Such β-strands could be considered as ‘assemblons’, independent associating units, by homology to the foldons (independent folding unit).
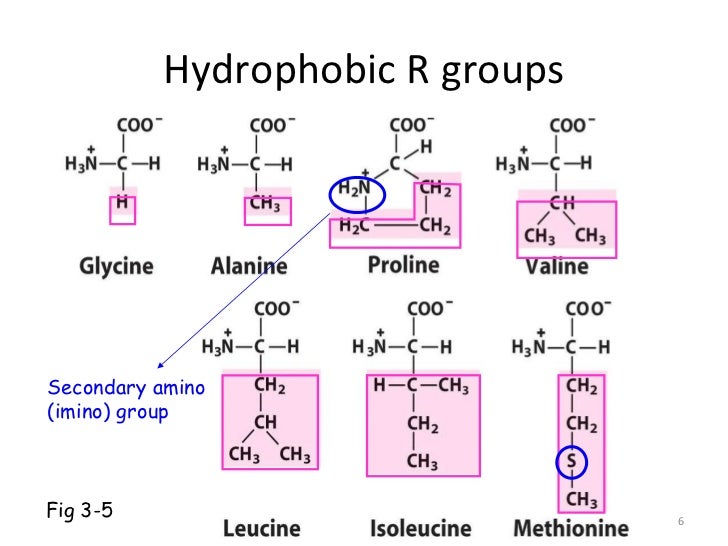
Thus, their sequences contain the features necessary for a β-interface formation. Moreover, the β-strands of the cholera toxin B subunit interface, when produced individually as synthetic peptides, are capable of inhibiting the assembly of the toxin into pentamers. This might open new venues for drug designs and predictive tool developments. Such charge distribution helps discriminating between sequences of intermolecular β-strands, of intramolecular β-strands and of β-strands forming β-amyloid fibers.

The intermolecular specificity is provided by the side chain networks via positioning different types of charged residues at the extremities (arginine) and in the middle (glutamic acid and histidine) of the interface. The secondary structure of the β-interfaces is implemented through the backbone networks which are enriched with the hydrophobic amino acids favored in intramolecular β-sheets (MCWIV). Each one has its own characteristics which can be associated to a distinct role. One of them involves interactions between main chain atoms (backbone network) while the other involves interactions between side chain and backbone atoms or between only side chain atoms (side chain network). The results show that the β-interfaces are made of two interdigitated interaction networks.
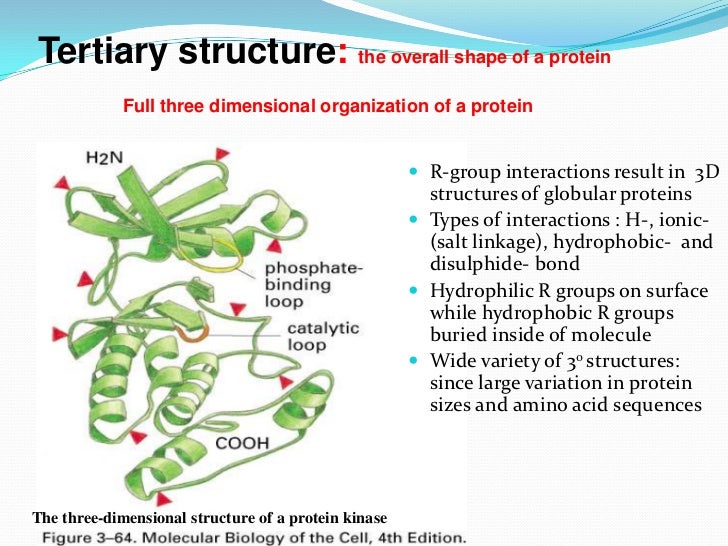
The protein oligomers of the dataset share the same geometry of interface, made by the association of two individual β-strands (β-interfaces), but are otherwise unrelated. The protein interfaces of 40 soluble protein oligomers of stœchiometries above two are investigated using a quantitative and qualitative methodology, which analyzes the x-ray structures of the protein oligomers and considers their interfaces as interaction networks.
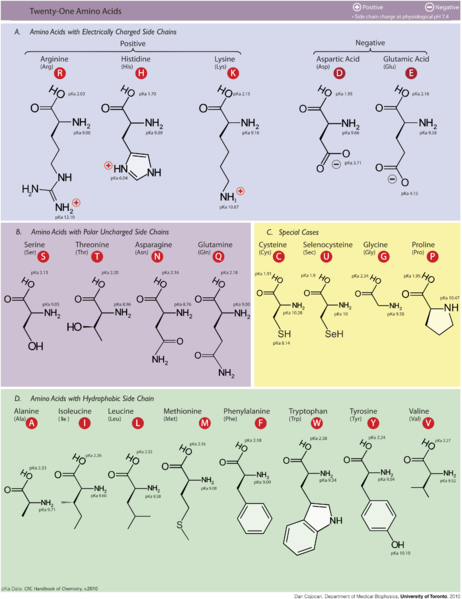
The protein chains are associated through intermolecular interactions constituting the protein interface. Protein oligomers are formed either permanently, transiently or even by default.
